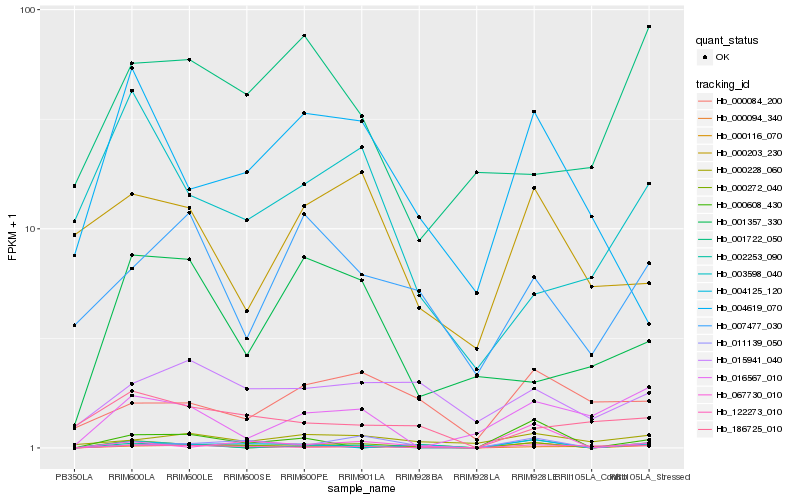
| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_000272_040 |
0.0 |
- |
- |
- |
| 2 |
Hb_000116_070 |
0.2775678582 |
transcription factor |
TF Family: C2H2 |
hypothetical protein JCGZ_22381 [Jatropha curcas] |
| 3 |
Hb_002253_090 |
0.2812252318 |
- |
- |
Chitinase 1 precursor, putative [Ricinus communis] |
| 4 |
Hb_011139_050 |
0.2869402141 |
- |
- |
nicotinate phosphoribosyltransferase family protein [Populus trichocarpa] |
| 5 |
Hb_122273_010 |
0.2881811018 |
- |
- |
DNA/RNA polymerases superfamily protein [Theobroma cacao] |
| 6 |
Hb_004125_120 |
0.2999391002 |
- |
- |
protein phosphatase 2c, putative [Ricinus communis] |
| 7 |
Hb_016567_010 |
0.3142255092 |
- |
- |
- |
| 8 |
Hb_003598_040 |
0.3213830538 |
transcription factor |
TF Family: HB |
PREDICTED: homeobox-leucine zipper protein ATHB-8 [Jatropha curcas] |
| 9 |
Hb_000228_060 |
0.3258275244 |
- |
- |
hypothetical protein JCGZ_21532 [Jatropha curcas] |
| 10 |
Hb_015941_040 |
0.3268644879 |
- |
- |
L-galactono-1,4-lactone dehydrogenase, mitochondrial [Cucumis sativus] |
| 11 |
Hb_000608_430 |
0.332199163 |
transcription factor |
TF Family: M-type |
mads box protein, putative [Ricinus communis] |
| 12 |
Hb_000094_340 |
0.3322286447 |
- |
- |
PREDICTED: LOW QUALITY PROTEIN: uncharacterized protein LOC103343662 [Prunus mume] |
| 13 |
Hb_004619_070 |
0.3325487685 |
- |
- |
hypothetical protein POPTR_0006s00550g [Populus trichocarpa] |
| 14 |
Hb_007477_030 |
0.3352754771 |
- |
- |
PREDICTED: monothiol glutaredoxin-S10-like [Jatropha curcas] |
| 15 |
Hb_000084_200 |
0.3400518268 |
- |
- |
PREDICTED: uncharacterized protein LOC105634688 isoform X2 [Jatropha curcas] |
| 16 |
Hb_186725_010 |
0.3410965215 |
- |
- |
PREDICTED: uncharacterized protein LOC105636491 [Jatropha curcas] |
| 17 |
Hb_001722_050 |
0.3419950965 |
- |
- |
calcium binding family protein [Populus trichocarpa] |
| 18 |
Hb_000203_230 |
0.3427589428 |
- |
- |
PREDICTED: probable inactive dual specificity protein phosphatase-like At4g18593 [Jatropha curcas] |
| 19 |
Hb_001357_330 |
0.3434556322 |
- |
- |
PREDICTED: paraspeckle component 1 [Jatropha curcas] |
| 20 |
Hb_067730_010 |
0.3441573104 |
- |
- |
hypothetical protein OsJ_21419 [Oryza sativa Japonica Group] |