About this database
Rubber Genome and Transcriptome Database
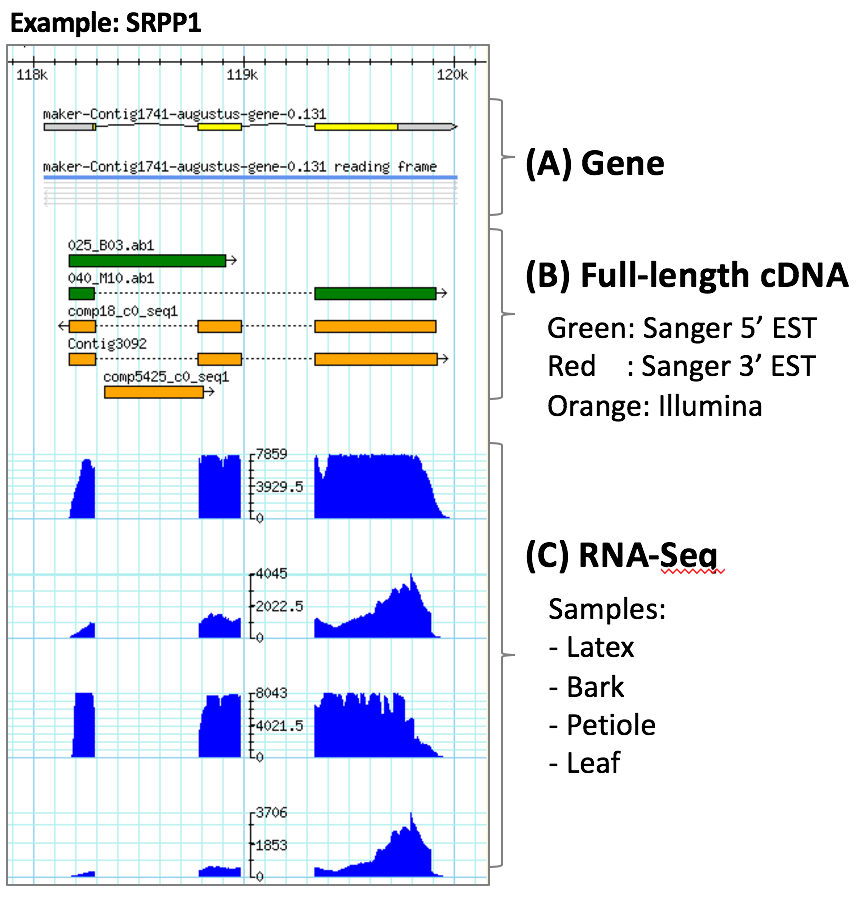
(A) Rubber draft genome was assembled based on ~155-fold combined coverage with Illumina and PacBio sequence data and has a total length of 1.55 Gb with 72.5% comprising repetitive DNA sequences. A total of 84,440 protein-coding genes were predicted and users can access these information on this database.
(B) As a next step, we constructed rubber full-length cDNA libraries sequencing around 20,000 clones by the Sanger method and over 15,000 contigs by Illumina sequencing. We annotated around 9,500 transcription start sites with Sanger 5’ ESTs.
(C) To elucidate the rubber biosynthetic pathways and their transcriptional regulators, we carried out tissue- and clone-specific RNA-Seq analysis. We compared RNA expression of leaves, petioles, bark and latex. As for clone-specificity, we extracted RNAs from latex in RRIM 600, RRIM 901 and PB 350 clones. In addition to our original data, we show publicly available RNA-Seq data.
Genome browser: http://matsui-lab.riken.jp/JBrowse/index.html?data=data/rubber
BLAST search: http://matsui-lab.riken.jp/rubber/blast/blast_rubber.html
Tutorial
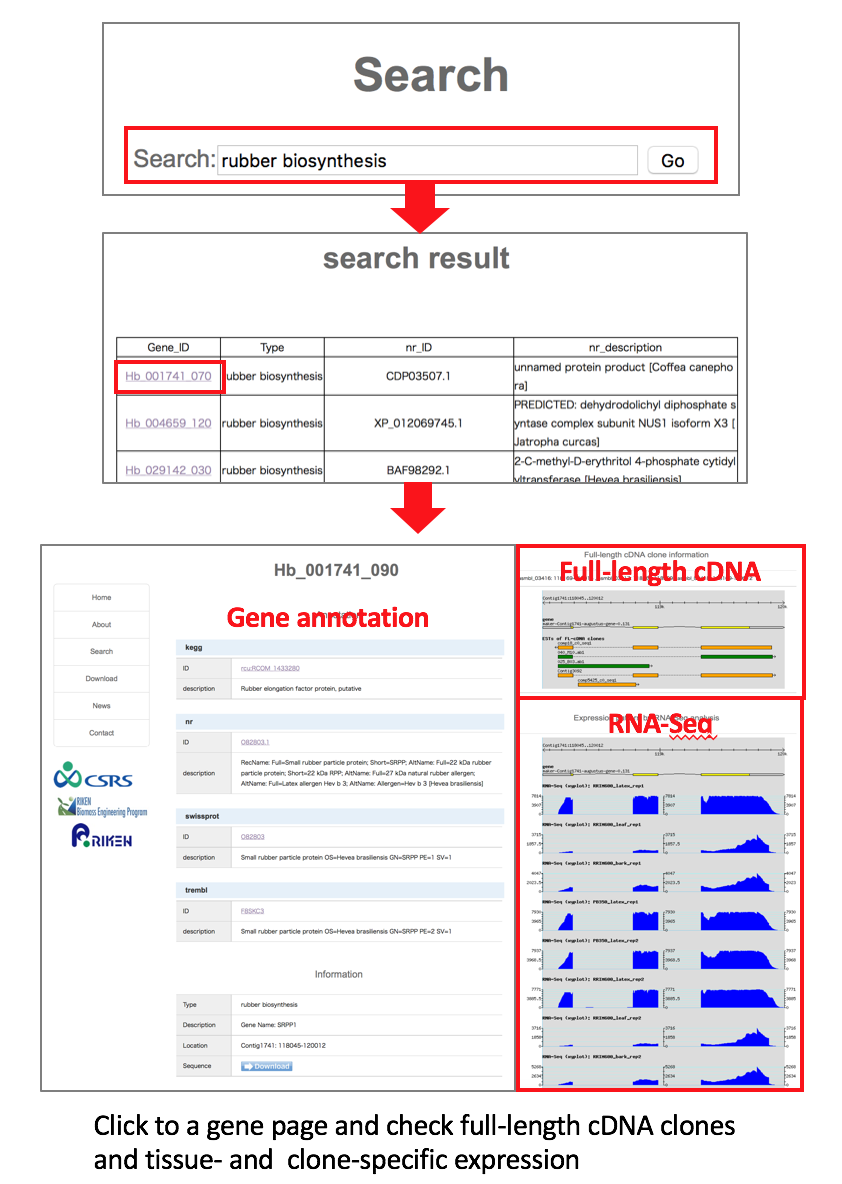
(1) Check the tissue specific expressions with keyword search.Go to Search page and type in any keyword. We used ”rubber biosynthesis” as an example.
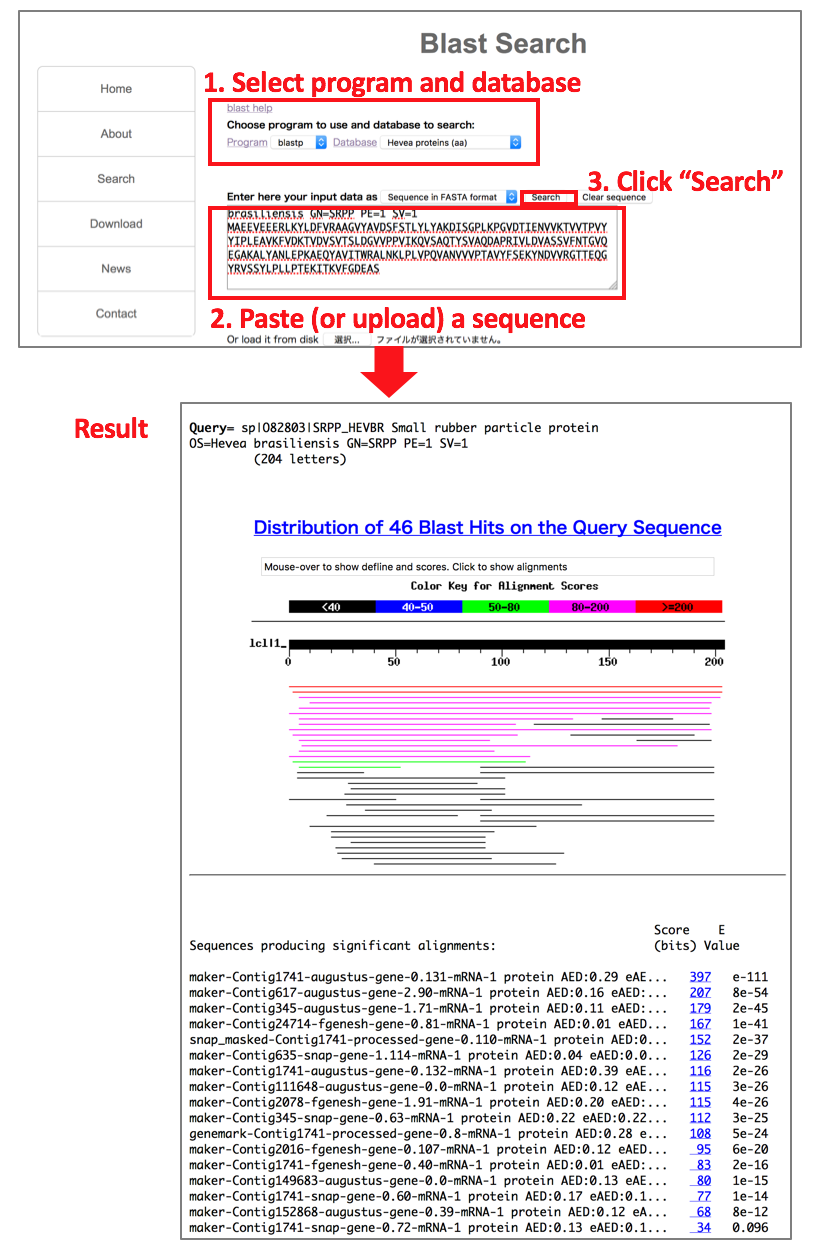
(2) Search genes by sequence homology (blast)
Go to blast search page: http://matsui-lab.riken.jp/rubber/blast/blast_rubber.html