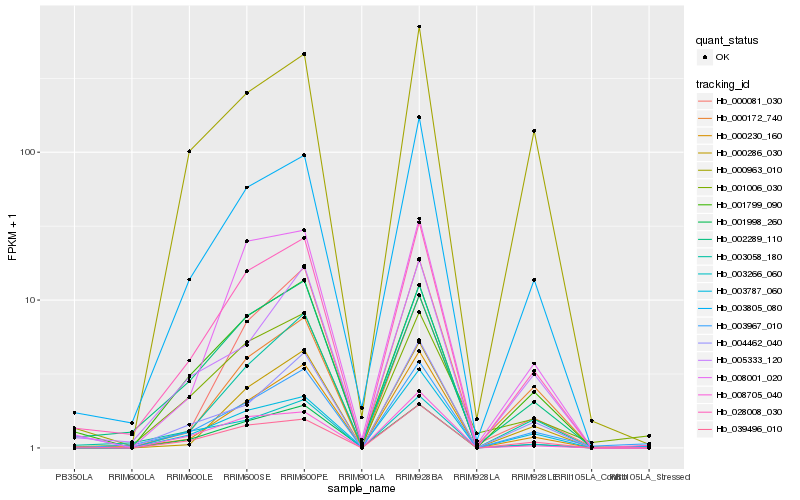
| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_028008_030 |
0.0 |
- |
- |
probable xyloglucan endotransglucosylase/hydrolase protein 23 precursor [Malus domestica] |
| 2 |
Hb_003805_080 |
0.0790272729 |
- |
- |
PREDICTED: isoflavone reductase-like protein [Jatropha curcas] |
| 3 |
Hb_000230_160 |
0.0858425329 |
- |
- |
PREDICTED: uncharacterized protein LOC105631858 isoform X1 [Jatropha curcas] |
| 4 |
Hb_001799_090 |
0.0940480839 |
- |
- |
PREDICTED: uclacyanin-2-like [Jatropha curcas] |
| 5 |
Hb_001998_260 |
0.0942933851 |
- |
- |
cytochrome P450, putative [Ricinus communis] |
| 6 |
Hb_039496_010 |
0.0966626961 |
- |
- |
PREDICTED: protein TRANSPARENT TESTA 12-like [Jatropha curcas] |
| 7 |
Hb_003967_010 |
0.106528579 |
- |
- |
cytochrome P450, putative [Ricinus communis] |
| 8 |
Hb_008001_020 |
0.1089206207 |
transcription factor |
TF Family: GRAS |
PREDICTED: protein SHORT-ROOT-like [Populus euphratica] |
| 9 |
Hb_002289_110 |
0.1135063508 |
- |
- |
PREDICTED: leucine-rich repeat receptor-like tyrosine-protein kinase At2g41820 [Jatropha curcas] |
| 10 |
Hb_003787_060 |
0.1149053997 |
- |
- |
PREDICTED: mechanosensitive ion channel protein 8-like [Jatropha curcas] |
| 11 |
Hb_008705_040 |
0.1153421159 |
- |
- |
PREDICTED: probable inactive leucine-rich repeat receptor-like protein kinase At1g66830 [Jatropha curcas] |
| 12 |
Hb_003058_180 |
0.1249660903 |
transcription factor |
TF Family: HSF |
hypothetical protein POPTR_0001s28040g [Populus trichocarpa] |
| 13 |
Hb_005333_120 |
0.1271108207 |
- |
- |
PREDICTED: protein ASPARTIC PROTEASE IN GUARD CELL 2 [Jatropha curcas] |
| 14 |
Hb_000286_030 |
0.127767384 |
- |
- |
PREDICTED: putative receptor-like protein kinase At3g47110 [Jatropha curcas] |
| 15 |
Hb_001006_030 |
0.1307029566 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 16 |
Hb_000081_030 |
0.1309449883 |
- |
- |
BnaC09g05810D [Brassica napus] |
| 17 |
Hb_000172_740 |
0.1326460509 |
- |
- |
HXXXD-type acyl-transferase family protein, putative [Theobroma cacao] |
| 18 |
Hb_003266_060 |
0.1330472173 |
- |
- |
PREDICTED: basic 7S globulin-like [Populus euphratica] |
| 19 |
Hb_000963_010 |
0.1348245887 |
- |
- |
PREDICTED: aquaporin PIP2-4 [Jatropha curcas] |
| 20 |
Hb_004462_040 |
0.1352907955 |
- |
- |
unnamed protein product [Vitis vinifera] |