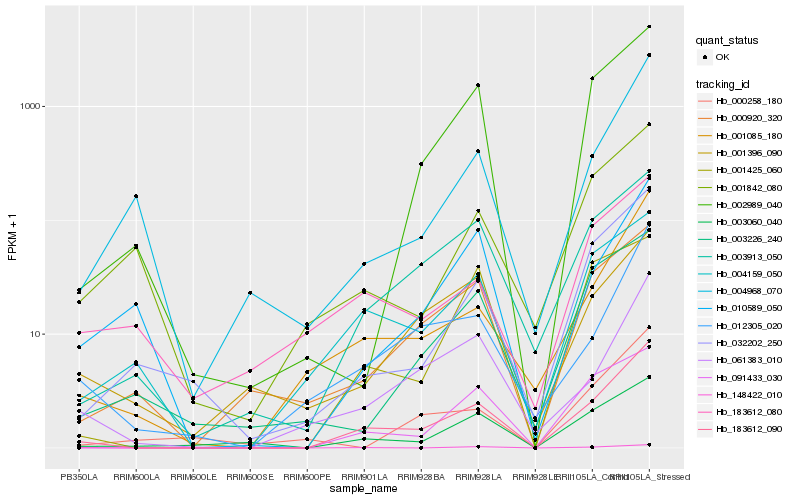
| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_183612_090 |
0.0 |
- |
- |
- |
| 2 |
Hb_001085_180 |
0.1566083743 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 3 |
Hb_012305_020 |
0.1733221445 |
- |
- |
beta-1,3-glucanase [Hevea brasiliensis] |
| 4 |
Hb_003060_040 |
0.1757639489 |
transcription factor |
TF Family: MIKC |
PREDICTED: agamous-like MADS-box protein AGL15 [Jatropha curcas] |
| 5 |
Hb_061383_010 |
0.1767573761 |
- |
- |
beta-1,3-glucanase [Hevea brasiliensis] |
| 6 |
Hb_003913_050 |
0.1793621061 |
- |
- |
60S ribosomal protein L27B [Hevea brasiliensis] |
| 7 |
Hb_091433_030 |
0.181020282 |
- |
- |
- |
| 8 |
Hb_004159_050 |
0.18419903 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 9 |
Hb_032202_250 |
0.1873811016 |
- |
- |
PREDICTED: F-box protein PP2-A13 [Jatropha curcas] |
| 10 |
Hb_000920_320 |
0.1892605 |
transcription factor |
TF Family: Orphans |
hypothetical protein JCGZ_24525 [Jatropha curcas] |
| 11 |
Hb_002989_040 |
0.1898319129 |
- |
- |
hypothetical protein 1 [Hevea brasiliensis] |
| 12 |
Hb_004968_070 |
0.2040345702 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 13 |
Hb_001396_090 |
0.2073375222 |
transcription factor |
TF Family: WRKY |
PREDICTED: probable WRKY transcription factor 75 [Jatropha curcas] |
| 14 |
Hb_010589_050 |
0.2074024218 |
- |
- |
- |
| 15 |
Hb_148422_010 |
0.2077840948 |
- |
- |
PREDICTED: uncharacterized protein LOC104908874, partial [Beta vulgaris subsp. vulgaris] |
| 16 |
Hb_183612_080 |
0.2152319445 |
- |
- |
hypothetical protein POPTR_0001s00500g [Populus trichocarpa] |
| 17 |
Hb_003226_240 |
0.2182313628 |
- |
- |
PREDICTED: caffeoylshikimate esterase [Jatropha curcas] |
| 18 |
Hb_001425_060 |
0.2223888906 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 19 |
Hb_001842_080 |
0.2242917796 |
- |
- |
- |
| 20 |
Hb_000258_180 |
0.2246218981 |
- |
- |
PREDICTED: glutathione S-transferase U10-like [Vitis vinifera] |