| Rank |
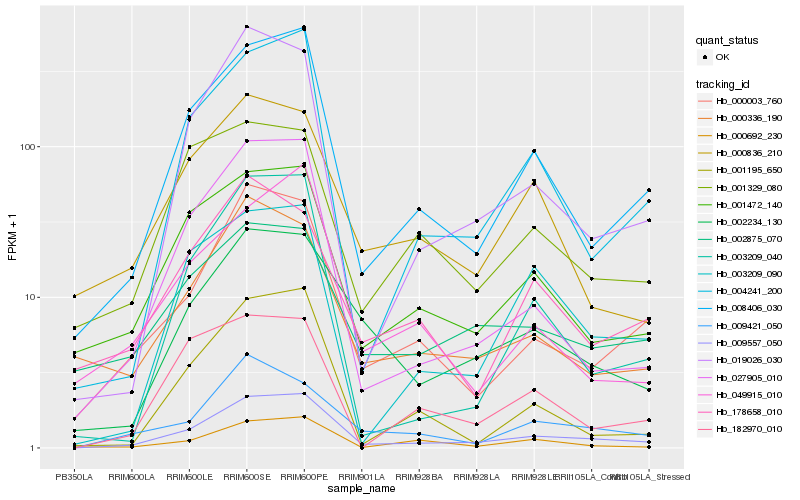
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
Hb_009557_050 |
0.0 |
transcription factor |
TF Family: C2H2 |
zinc finger family protein [Populus trichocarpa] |
| 2 |
Hb_008406_030 |
0.1114654957 |
transcription factor |
TF Family: ERF |
AP2/ERF super family protein [Hevea brasiliensis] |
| 3 |
Hb_004241_200 |
0.1415495702 |
- |
- |
hypothetical protein JCGZ_00744 [Jatropha curcas] |
| 4 |
Hb_000003_760 |
0.1424709919 |
transcription factor |
TF Family: AUX/IAA |
PREDICTED: auxin-responsive protein IAA26-like [Jatropha curcas] |
| 5 |
Hb_001472_140 |
0.1467421231 |
- |
- |
PREDICTED: BTB/POZ domain-containing protein At5g41330 [Jatropha curcas] |
| 6 |
Hb_002234_130 |
0.1484870528 |
- |
- |
ATPP2-A13, putative [Ricinus communis] |
| 7 |
Hb_019026_030 |
0.1532523499 |
- |
- |
hypothetical protein JCGZ_00744 [Jatropha curcas] |
| 8 |
Hb_027905_010 |
0.1597476408 |
- |
- |
transferase, transferring glycosyl groups, putative [Ricinus communis] |
| 9 |
Hb_003209_090 |
0.1660642022 |
- |
- |
Xyloglucan endotransglucosylase/hydrolase family protein [Theobroma cacao] |
| 10 |
Hb_000336_190 |
0.1698689565 |
- |
- |
PREDICTED: UPF0481 protein At3g47200-like [Gossypium raimondii] |
| 11 |
Hb_009421_050 |
0.1707107178 |
- |
- |
PREDICTED: uncharacterized protein LOC105647037 [Jatropha curcas] |
| 12 |
Hb_002875_070 |
0.1718445924 |
- |
- |
PREDICTED: probable mitochondrial chaperone BCS1-A [Jatropha curcas] |
| 13 |
Hb_003209_040 |
0.1719653673 |
- |
- |
PREDICTED: xyloglucan endotransglucosylase/hydrolase protein 22-like [Gossypium raimondii] |
| 14 |
Hb_000692_230 |
0.1759661906 |
- |
- |
phospholipase d delta, putative [Ricinus communis] |
| 15 |
Hb_001195_650 |
0.1787235365 |
- |
- |
PREDICTED: calcium uniporter protein 4, mitochondrial [Jatropha curcas] |
| 16 |
Hb_000836_210 |
0.1820223673 |
- |
- |
hypothetical protein JCGZ_14362 [Jatropha curcas] |
| 17 |
Hb_182970_010 |
0.1843544403 |
- |
- |
calmodulin-binding protein 60-D [Populus trichocarpa] |
| 18 |
Hb_049915_010 |
0.1857556853 |
- |
- |
PREDICTED: uncharacterized serine-rich protein C215.13 [Jatropha curcas] |
| 19 |
Hb_178658_010 |
0.1859708638 |
- |
- |
PREDICTED: uncharacterized protein LOC105641396 [Jatropha curcas] |
| 20 |
Hb_001329_080 |
0.1875630495 |
- |
- |
PREDICTED: probable protein phosphatase 2C 49 [Jatropha curcas] |