| Rank |
Gene |
Score (JSD) |
Function |
Description |
NCBI(nr) information |
| 1 |
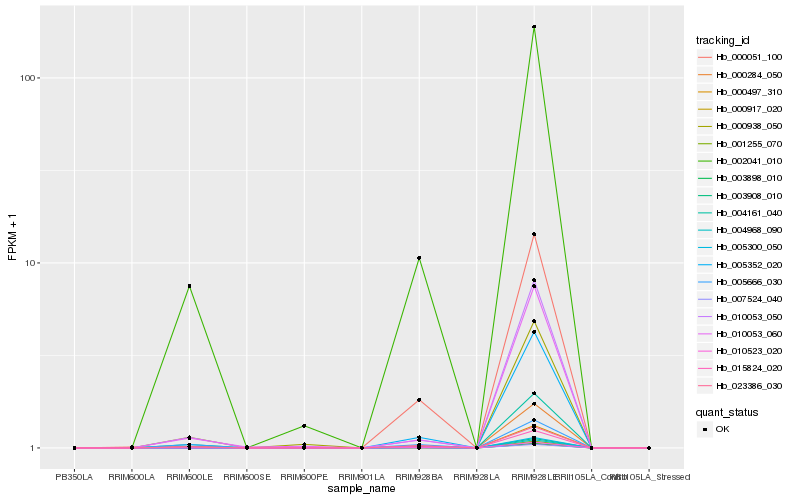
Hb_002041_010 |
0.0 |
- |
- |
- |
| 2 |
Hb_000497_310 |
0.0320986747 |
- |
- |
conserved hypothetical protein [Ricinus communis] |
| 3 |
Hb_000284_050 |
0.0635134613 |
- |
- |
hypothetical protein L484_026849 [Morus notabilis] |
| 4 |
Hb_015824_020 |
0.0774016976 |
- |
- |
- |
| 5 |
Hb_023386_030 |
0.0774345151 |
- |
- |
- |
| 6 |
Hb_010053_050 |
0.0904369454 |
- |
- |
hypothetical protein POPTR_0530s002102g, partial [Populus trichocarpa] |
| 7 |
Hb_010053_060 |
0.0921556363 |
- |
- |
ATP-binding cassette transporter, putative [Ricinus communis] |
| 8 |
Hb_004161_040 |
0.1130001751 |
- |
- |
PREDICTED: putative receptor-like protein kinase At4g00960 [Populus euphratica] |
| 9 |
Hb_004968_090 |
0.1229768969 |
- |
- |
PREDICTED: uncharacterized protein LOC104882781 [Vitis vinifera] |
| 10 |
Hb_000938_050 |
0.1244849671 |
- |
- |
hypothetical protein JCGZ_09640 [Jatropha curcas] |
| 11 |
Hb_005666_030 |
0.1244882155 |
- |
- |
PREDICTED: uncharacterized protein LOC103489421 [Cucumis melo] |
| 12 |
Hb_005300_050 |
0.130024602 |
- |
- |
hypothetical protein VITISV_001308 [Vitis vinifera] |
| 13 |
Hb_003908_010 |
0.1300418557 |
- |
- |
polyprotein [Oryza australiensis] |
| 14 |
Hb_001255_070 |
0.130090665 |
- |
- |
PREDICTED: LOW QUALITY PROTEIN: uncharacterized protein LOC104235718, partial [Nicotiana sylvestris] |
| 15 |
Hb_000917_020 |
0.1301360354 |
- |
- |
polyprotein [Citrus endogenous pararetrovirus] |
| 16 |
Hb_007524_040 |
0.1301486457 |
- |
- |
PREDICTED: LOW QUALITY PROTEIN: uncharacterized protein LOC101511096 [Cicer arietinum] |
| 17 |
Hb_005352_020 |
0.1306183023 |
- |
- |
- |
| 18 |
Hb_000051_100 |
0.1309383425 |
- |
- |
hypothetical protein MIMGU_mgv1a025286mg [Erythranthe guttata] |
| 19 |
Hb_003898_010 |
0.1309997068 |
- |
- |
hypothetical protein PRUPE_ppa022673mg [Prunus persica] |
| 20 |
Hb_010523_020 |
0.1310816875 |
- |
- |
DNA/RNA polymerases superfamily protein [Theobroma cacao] |